Operating system: macos
Slicer version:4.13
Expected behavior: change nii segmentation to .vtk
Actual behavior: the vtk format is not available for export mesh
I have a binary 3d nii image and I am trying to make a mesh of it and save it in .vtk directly using slicer software preferably.
the procedure I tried based on the tutorial of slicer, I opened my image as segmentation, then I went to segmentation modules and made the closed surface by clicking on the closed surface representations. Then in the export part I chose operation: export, output type: model. In the export to file section I chose a directory to save but there is no vtk format there for save it in vtk. I would be very grateful if you could help in that.
Thanks, That works ! I am actually making the mesh and save it to vtk in order to open it in paraview and resample the p-value surface on it but it seems that after saving in vtk, some specifications is changed since it does not overlay my other surface when opening the vtk image in paraview. How can I save .vtk in the same origin or change the origin of it after saving?
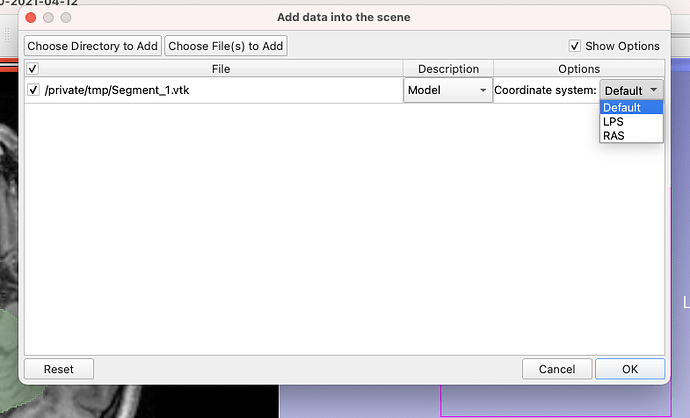
Thank you, based on the image in the previous reply, I opened it as RAS and then saved it in vtk (poly data) and then again opened it in paraviwe but it did not change and still is exactly similar to one before:
i have attached a photo here when I opened it in paraview , my mesh is the blue one but it should be in the place of the white one and flipped as the white (correct) one is.
Could you please explain a bit more about how to harden it through a transform before I save?
the segment image actually is aligned correctly before I save as vtk but the vtk file is not correctly placed and aligned with my probability surface map after conversion.
You need to create a linear transform in the Transforms module and change the first two 1s on the main diagonal to be -1. Then you apply that transform to the model (select the model click the green arrow at the bottom of the Transforms module). Then click the harden transform button. HTH

Paraview’s volume support is very limited. For example, it completely ignores image axis directions. Slicer display, volume rendering GUI is very poor, there is barely any support transforms, etc. For this reason, it is very inconvenient to use it in any serious medical image computing projects. Image orientation support has been recently added to VTK, so at some point Paraview might implement support for oriented images.
I would recommend to copy p values from the volume to the segmented surface using “Probe volumes with model” module in Slicer.
thanks for your reply but it actually when i save it after transformation it does not change the orientation or anything else it is exactly the same as the image before. As you said I loaded the previously saved vtk image as RAS and in the transformation module I changed the two 1s to -1 and then select the image (segment_1_main) on the left as shown below and click on the first arrow to bring it to the right and clicked on the hard transform then from the save option above I saved it as .vtk again. are all process correct?
VTK format cannot save the image orientation. We generally recommend nrrd format for saving 3D volumes. For simple things metaimage works well, too. For neuroimaging you may choose nifti.
Thanks for your reply. Actually I am using vtk or vtp because I am going to use that in paraview which accepts the vtk and vtp formats. There should be a way since the white mesh image in the few images before I took screenshot from paraview shows the correct mesh which has been made from the same segment binary image so I think there is a way to make that correction in the mesh i have saved with slicer. Actually I have been informed that the white mesh has also been made with slicer. So, I am trying slicer. So, I hope I could make my blue mesh correct as the white one is.
Paraview can read uncompressed metaimage and nrrd formats, too. But it does not help much, because Paraview ignores the image orientation information.
I see that you are referring to VTP. Maybe you mean you want to save meshes and not images? Meshes can be saved in any orientation and Paraview will load them correctly.
Paraview may still show misaligned image and mesh because it cannot load the image correctly. The only way you can work around the Paraview’s non-oriented image issue is that if you transform everything (the image, segmentation, all meshes) with a transformation that makes the image’s direction matrix become identity (diagonal of 1.0), because it does not matter if you ignore an identity orientation matrix. You need to harden this transform on all nodes before saving. I’m not sure if you need to save the meshes in LPS and RAS coordinate system in this case, so try both.
Paraview is great for working with meshes, but if you work with image volumes then you are much better off using Slicer. Let us know if there is anything that you know how to do in Paraview but not sure how to do in Slicer. Slicer is built on top of VTK the same way as Paraview, so Slicer has the same low-level features, just not all have been made available via graphical user interface or not the same way as in Paraview.