I’ve been using the ANTs extension for registering two single-label volumes to eachother with good results. In an attempt to improve registration robustness, I started incorporating fixed and moving masks (FM, MM), but this is yielding some unexpected results. To simplify the issue I’m just focusing on a single stage affine transform. The affine transform with the fixed mask alone looks very reasonable; but if I add a moving mask to this same affine configuration (along with the fixed) the registration is much poorer. For now, the moving mask is essentially just a bounding box with zeros at the edges.
When working properly, the largest components of the final affine transform are a Z translation (shift) and a Z scaling (expansion). Something about showing the MM (in addition to FM) loses this info. As a test, I ran a “good” FM only registration but also included a 2nd dummy affine stage (following the correct/complete one), where there are zero convergence steps, but now both the FM and MM are shown. This somehow changes the 3-3 matrix value (Z scaling factor) from 1.2 (ideal) to ~1. If this dummy step only has the FM, the “affine erasure” doesn’t happen.
Trouble shooting I created a MM that is “all 1s”, expecting that it should work the same as a having no moving mask but this wasn’t the case, the all 1s MM worsened the quality.
Any chance this is a bug? Might the image origin of the moving mask be getting read improperly? Perhaps I’m just ignorant of how masks interact by design in ANTs.
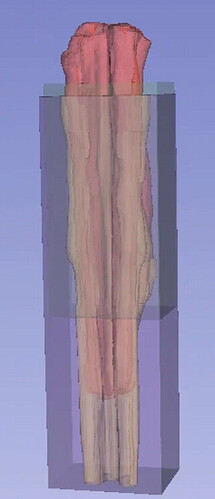
If it’s any help I’m including an image
Red: Fixed volume (my tissue); Yellow: Moving volume (atlas); Green: fixed mask; Purple: moving mask
**Note, this doesn’t show the dimensions of the zero values. For example, how large the virtual space of the fixed volume is.