Hi,
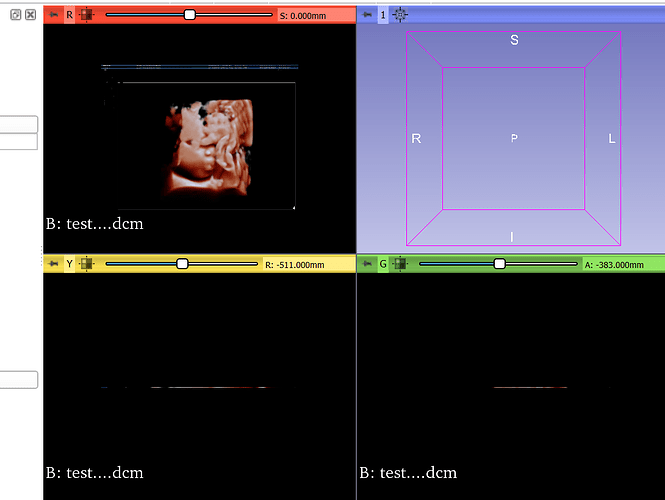
I’ve received a .dcm file that only displays on the axial view as a 2D image of a sonogram. Is it possible to convert this into a 3D volume for segmentation? I’ve tried increasing the volumes using the Volumes module, as well as convert to DICOM module but I get the same thing. If not, what type of file is preferred in 3DSlicer when wanting to segment fetal ultrasounds/sonograms. Attached is the screenshot of what I get when I uploaded the .dcm file onto 3DSlicer.
This seems like a Secondary Capture, basically a 2D screenshot. This is a completely valid DICOM format, one of many types.
Do you have more files? If not, then it is not possible to have a 3D volume. Such volume renderings are usually done in the ultrasound workstation and saved as image. Real reconstructed 3D ultrasound volumes are very rare in current clinical settings.
It is possible that standard DICOM fields only contain this 2D screenshot but there is a full 3D volume stored in private fields. SlicerHeart extension can load many kinds of 3D/4D volumes.
Thank you for your response! Do you know how (or where I can learn) to access/edit the private fields?
You can view content of all DICOM fields in DICOM module (right-click on a series and choose “View DICOM metadata”). However, 3D ultrasound data is usually stored using complex encoding so you cannot trivially get the 3D volume out of it. That’s what SlicerHeart extension takes care of, for a number of ultrasound systems. See DICOM import section for more details.
If it is not clear enough how you can get the 3D volume then give us more information (brand and model of the ultrasound system, file size, maybe the whole DICOM metadata - with patient information excluded) and we can provide more specific advice.
I understand. I will take a look at the section, and see if i can also get any more information to give you. Thanks again.
hi, this seems to be a constant topic for … most of us.
how to get the geometry from a DICOM file that seems to have only a screenshot.
can you help me?
i have a Mindray DC40 and a Mindray Renewa i9.
The new Ultrasound extension (that will be officially announced soon but it is already available in the Extensions Manager) should retrieve all geometry data that is available in the DICOM header. @ungi, developer of this new extension may provide more details if you tell a bit more about your project, what your end goal is, and how would you use the ultrasound images.
Hi @fili_S_S, have you tried to load you DICOM files in a recent preview version of Slicer? @lassoan recently updated how time series (ultrasound) are loaded in Slicer. I tested it with Mindray DICOM files and the pixel spacing was correct.
If you need the full geometry of the ultrasound (e.g. four corners of a fan shaped image), that is not stored in DICOM unfortunately. We needed those points and I added a module in the Ultrasound extension to manually annotate the corner points and save them in separate files besides the DICOMs.
There are other confusing features of Minday DICOM files. E.g., if you don’t change the imaging parameters on the machine, it stops saving image geometry in the DICOM headers. Instead, it keeps the same series ID, so all scans are loaded at the same time, and image geometry is loaded from the first file. That is bad because we cannot throw out scans that we don’t need for a project. I’m planning to address these issues as much as possible in the Ultrasound extension, but it is a work-in-progress.
Basically I’m trying to 3d print fetal heads for NON COMERCIAL uses (gifts for patients in my private practice). I’m trying go obtain fetus head models or early pregnancy embryo models