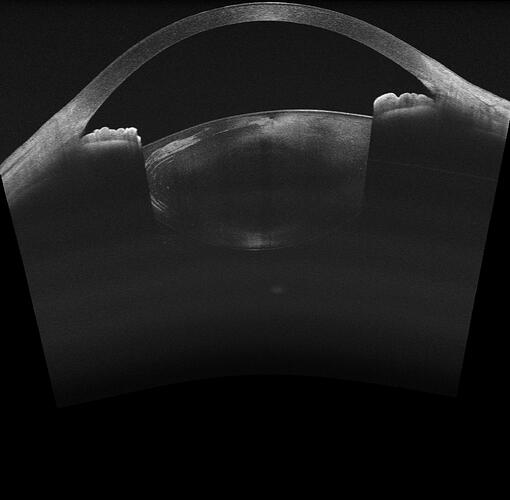
the jpg like this
Yes, sure, you can render even 16 slices in 3D. Are the slices parallel or rotated?
Rotated,and I attempted to reconstruct based on the previous posts about OCT image reconstruction, but I’m not clear about the functionality of the extension modules. Should I use IGT or slicerhreat? If possible, could you provide a more detailed explanation? Thank you very much!
16 slices for 180 degrees would be quite sparse (you have very high resolution slices with huge gaps between them), but depending on what you want to do, you might get a usable result.
SlicerHeart Reconstruct4DCineMRI module can serve as a good example of how to organize image a set of slices into a time sequence of 3D volumes, but since you only have a single timepoint, there is not a lot to use from there.
Instead, you can use this code snippet to put together a sequence of image slices, with each slice having a correct position and orientation and then reconstruct a volume from that.
Since your data is extremely sparse. you would end up with big gaps between them, which could make it difficult to interpret the 3D volume. You could fix that by adding interpolated slices (compute weighted average of neighbor slices and insert them between your original slices).
I used the code snippet you provided, and when I reached the model reconstruction step, the CPU usage significantly increased, and then the slicer crashed. I tried multiple times, but even when it completed running, I couldn’t obtain the correct model. When importing JPG images, should I just import them from the folder to form a single model volume, or should I import them separately multiple times to get multiple model volumes?
This sounds promising. The application crashed because it has ran out of memory. The volume reconstructor automatically allocates a memory buffer that is large enough to contain all the slices at the desired resolution, and if the the slices cover a large region and the spacing you have chosen for the reconstructed volume is small then this memory buffer will be larger than the amount of memory your computer has. The problem is either in how you set the image positions/orientations (you can verify this by browsing the generated image sequence using Sequence browser toolbar and show the slice positions by configuring Volume Reslice Driver module) or choose a larger output spacing (to begin with, use 10x or 100x larger output spacing then the spacing of your input image; if volume reconstruction is successful you can gradually decrease this value to see what is the smallest spacing your computer can handle).
The result of volume reconstruction is not a model (i.e., mesh), but an image (3D volume). You can see it in slice views or use Volume rendering module to show it in 3D.
The code snippet I linked is for getting slices from a single volume, so probably the easiest is to load the JPG stack as a single volume.