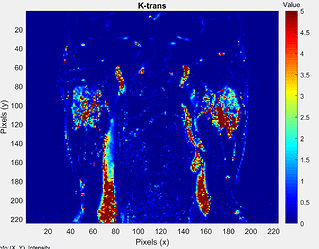
Is there a way to check if you are losing data between Slicer and MATLAB? I found an alternative method using advice given from the Normalization/calibration - #5 by fedorov thread , and in particular using the ShiftScaleImageFilter on each of the maps. Since now I know DICOM format only accepts integer values on Slicer, then by scaling each map by 1000 (or to whichever accuracy needed) prior to export, then you can keep the decimal values and just re-scale back within MATLAB easy enough (see image below). Not sure about data-loss in this process though.
madeline
(Madeline Carr)
7
Related topics
| Topic | Replies | Views | Activity | |
|---|---|---|---|---|
| DICOM file from Slicer viewed differently in Materialise Mimics | 22 | 1934 | November 25, 2020 | |
| PkModeling tutorial | 3 | 814 | November 21, 2019 | |
| Issue on exporting DICOM images | 4 | 494 | August 13, 2020 | |
| Conversion of FA (Fractional Anisotropy) map to DICOM failing | 5 | 267 | December 28, 2023 | |
| Saving dicom to dicom, the exported and input do not match | 9 | 536 | April 28, 2022 |